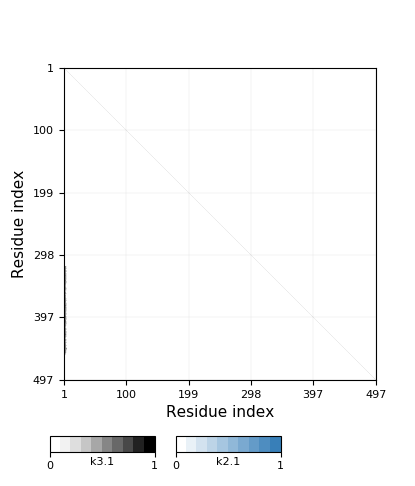
|
|
|
|
K -31
|
Chain closureGlu2 <-> Glu498 ... His461 <-> Bridging ionZn504 <-> Asp269 ... Asp311 <-> Bridging ionZn505 <-> Ser65 ... Glu2 |
probabilistic |
|
|
|
|
|
K -31
|
Chain closureGlu2 <-> Glu498 ... His461 <-> Bridging ionZn504 <-> Asp269 ... His312 <-> Bridging ionZn505 <-> Ser65 ... Glu2 |
probabilistic |
|
|
|
|
|
K -31
|
Chain closureGlu2 <-> Glu498 ... His461 <-> Bridging ionZn504 <-> His273 ... Asp311 <-> Bridging ionZn505 <-> Ser65 ... Glu2 |
probabilistic |
|
|
|
|
|
K -31
|
Chain closureGlu2 <-> Glu498 ... His461 <-> Bridging ionZn504 <-> His273 ... His312 <-> Bridging ionZn505 <-> Ser65 ... Glu2 |
probabilistic |
|
|
|
|
|
K -31
|
Chain closureGlu2 <-> Glu498 ... His461 <-> Bridging ionZn504 <-> Asp269 ... Asp311 <-> Bridging ionZn505 <-> Ser65 ... Gly48 <-> Bridging ionMg502 <-> Ala45 ... Glu2 |
probabilistic |
|
|
|
|
|
K -31
|
Chain closureGlu2 <-> Glu498 ... His461 <-> Bridging ionZn504 <-> Asp269 ... Asp311 <-> Bridging ionZn505 <-> Ser65 ... Gly48 <-> Bridging ionMg502 <-> Lys46 ... Glu2 |
probabilistic |
|
|
|
|
|
K -31
|
Chain closureGlu2 <-> Glu498 ... His461 <-> Bridging ionZn504 <-> Asp269 ... His312 <-> Bridging ionZn505 <-> Ser65 ... Gly48 <-> Bridging ionMg502 <-> Ala45 ... Glu2 |
probabilistic |
|
|
|
|
|
K -31
|
Chain closureGlu2 <-> Glu498 ... His461 <-> Bridging ionZn504 <-> Asp269 ... His312 <-> Bridging ionZn505 <-> Ser65 ... Gly48 <-> Bridging ionMg502 <-> Lys46 ... Glu2 |
probabilistic |
|
|
|
|
|
K -31
|
Chain closureGlu2 <-> Glu498 ... His461 <-> Bridging ionZn504 <-> His273 ... Asp311 <-> Bridging ionZn505 <-> Ser65 ... Gly48 <-> Bridging ionMg502 <-> Ala45 ... Glu2 |
probabilistic |
|
|
|
|
|
K -31
|
Chain closureGlu2 <-> Glu498 ... His461 <-> Bridging ionZn504 <-> His273 ... Asp311 <-> Bridging ionZn505 <-> Ser65 ... Gly48 <-> Bridging ionMg502 <-> Lys46 ... Glu2 |
probabilistic |
|
|
|
|
|
K -31
|
Chain closureGlu2 <-> Glu498 ... His461 <-> Bridging ionZn504 <-> His273 ... His312 <-> Bridging ionZn505 <-> Ser65 ... Gly48 <-> Bridging ionMg502 <-> Ala45 ... Glu2 |
probabilistic |
|
|
|
|
|
K -31
|
Chain closureGlu2 <-> Glu498 ... His461 <-> Bridging ionZn504 <-> His273 ... His312 <-> Bridging ionZn505 <-> Ser65 ... Gly48 <-> Bridging ionMg502 <-> Lys46 ... Glu2 |
probabilistic |
|

 Genus: 186
Genus: 186